Spicy new research: Chili aptamer structure solved!
10.06.2021The structure of the fluorogenic Chili RNA aptamer and its mechanism of fluorescence activation by excited state proton transfer to the RNA are now reported in Nature Communications.
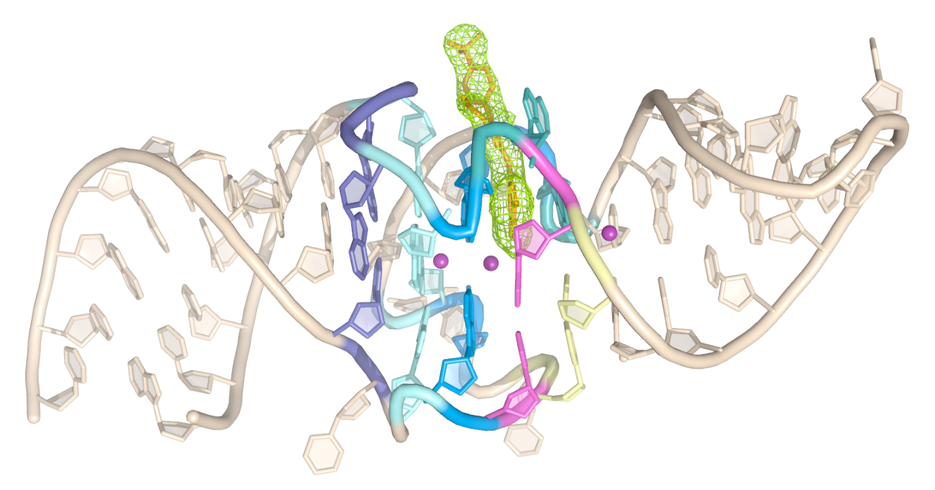
The Chili RNA aptamer mimics large Stokes shift fluorescent proteins. The 52-nt RNA specifically binds and activates conditional fluorophores to elicit fluorescence emission. Two co-crystal structures structures of the Chili aptamer in complex with the green-emitting DMHBI+ and the orange-red-emitting DMHBO+ ligands revealed a G-quadruplex and a trans-sugar-sugar edge G:G base pair that immobilize the ligand by pi-pi stacking. A Watson-Crick G:C base pair in the fluorophore binding site establishes a short hydrogen bond between the N7 of guanine and the phenolic OH of the ligand. The Hoogsteen side of this GC base pair on top of a guanine quartet together with a potassium ion establisheds the structural motif that facilitates ultrafast excited state proton transfer (ESPT) on the 100 fs timescale. Atomic mutagenesis confirmed the mode of action of the large Stokes shift fluorogenic RNA aptamer. These results strengthen the view of G-quadruplexes flanked by canonical A-form duplexes as versatile and privileged architectures for fluorescence activation of HBI chromophores.
see: Nat. Commun. 2021, 12. First published 10 June 2021.