Deoxyribozymes
Artificial evolution of catalytic DNA
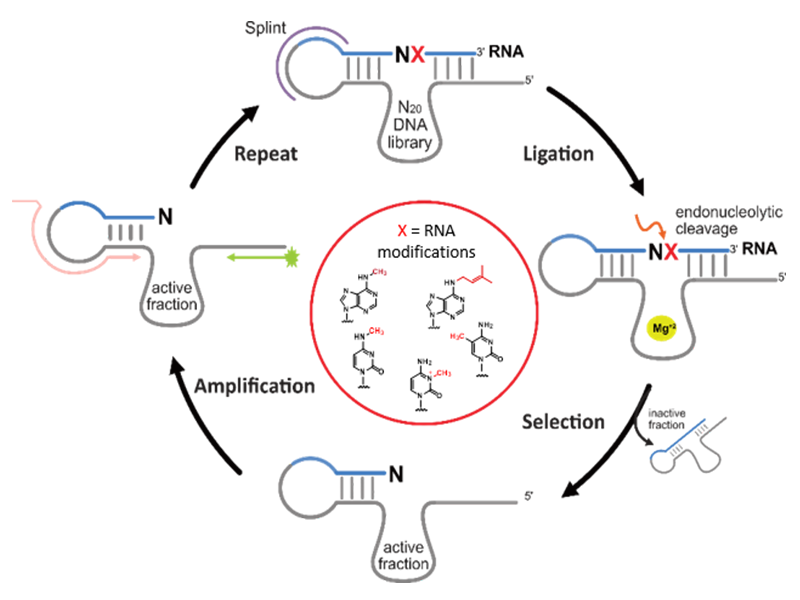
Using in vitro selection, we developed specific RNA‐cleaving deoxyribozymes for the detection of RNA modifications and to interrogate the methylation state of target RNAs, providing an alternative method for the biochemical validation of RNA modification sites. We established DNAzymes for analysis of N6‐methyladenosine (m6A) and i6A in various types of RNA, and recently found deoxyribozymes that distinguish the three natural methylated cytidine isomers m3C, m4C and m5C in synthetic as well as native human mitochondrial tRNAs.
Publications on this topic:
M.V. Sednev, V. Mykhailiuk, P. Choudhury, J. Halang, K.E. Sloan, M.T. Bohnsack, C. Höbartner,
N6‐Methyladenosine‐Sensitive RNA‐Cleaving Deoxyribozymes
Angew. Chem. Int. Ed. 2018, 57, 15117-15121. (VIP) Angew. Chem. 2018, 130, 15337-15341
A. Liaqat, C. Stiller, M. Michel, M.V. Sednev, C. Höbartner
N6-isopentenyladenosine in RNA determines the cleavage site of endonuclease deoxyribozymes
Angew. Chem. Int. Ed. 2020, 59, 18627-18631.
A. Liaqat, M.V. Sednev, C. Stiller, C. Höbartner
RNA-cleaving deoxyribozymes differentiate methylated cytidine isomers in RNA
Angew. Chem. Int. Ed. 2021, 60, 19058–19062.
High-throughput activity profiling
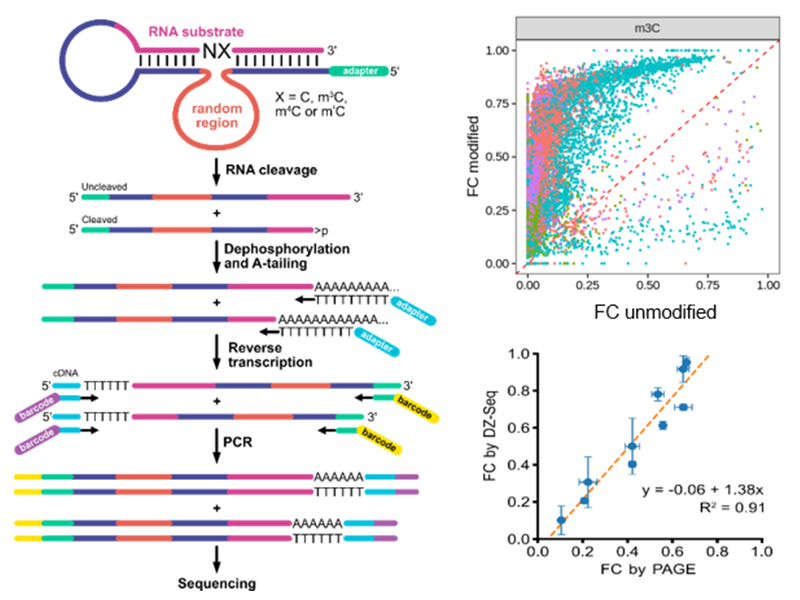
Building on in vitro selection of modification-specific deoxribozymes, we developed a high-throughput sequencing method, named DZ-seq, which directly measures activity and localizes cleavage sites of thousands of deoxyribozymes in parallel. DZ-seq exploits A-tailing followed by reverse transcription with an oligo-dT primer to capture cleavage status and sequences of both deoxyribozyme and RNA substrate. We validated DZ-seq by conventional analytical methods and demonstrated its utility by discovery of novel deoxyribozymes. The method has high potential for the discovery of additional modification-specific deoxyribozymes.
Publications on this topic:
M. V. Sednev, A. Liaqat, C. Höbartner
High-Throughput Activity Profiling of RNA-Cleaving DNA Catalysts by Deoxyribozyme Sequencing (DZ-seq)
J. Am. Chem. Soc. 2022 144, 2090-2094.
Structure of DNA catalyst
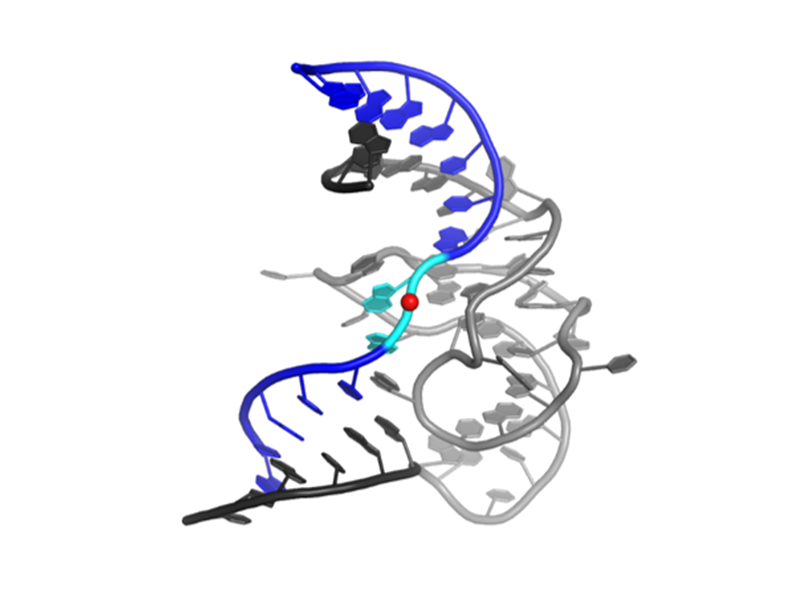
Despite more than two decades of biochemical study, mechanistic understanding of deoxyribozymes has been limited by the lack of any high-resolution structures of a deoxyribozyme until 2016, when we first presented the 2.8-Å-resolution crystal structure of the RNA-ligating deoxyribozyme 9DB1 in the postcatalytic state.
Compared to ribozymes, the sugar-phosphate backbone of the catalytic domain of the DNA enzyme 9DB1 has greater conformational diversity, thus compensating for the lack of the 2′-hydroxyl group. The structure confirms the idea that DNA can form complex tertiary folds that support catalysis.
A. Ponce-Salvatierra, K. Wawrzyniak-Turek, U. Steuerwald, C. Höbartner, V. Pena, Crystal structure of a DNA catalyst.
Nature 2016, 529, 231-234
C. Höbartner, How DNA catalyzes RNA ligation
Nature Catalysis 2019, 2, 483–484.
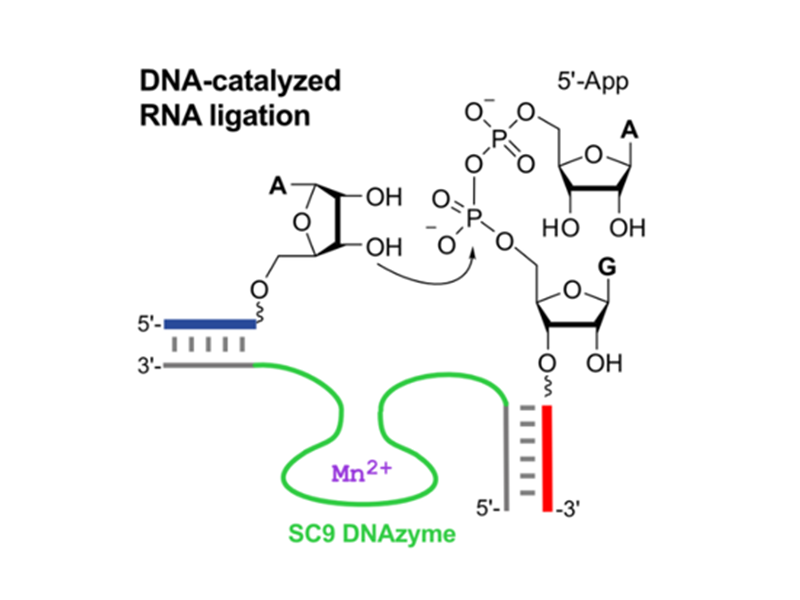
DNA-catalyzed RNA ligation
A novel class of RNA ligase deoxyribozymes was found by in vitro selection. These DNAzymes use 5′-adenylated RNA (5′-AppRNA) as the donor substrate, mimicking the activated intermediates of protein-catalyzed RNA ligation to catalyze the intermolecular linear RNA-RNA ligation via the formation of a native 3′-5′-phosphodiester linkage. These tools allow the protein-free synthesis of long RNAs, for example containing precious modified nucleotides or fluorescent labels for biochemical and biophysical investigations.
C.P.M. Scheitl, S. Lange, C. Höbartner
New Deoxyribozymes for the Native Ligation of RNA
Molecules 2020, 25, 3650